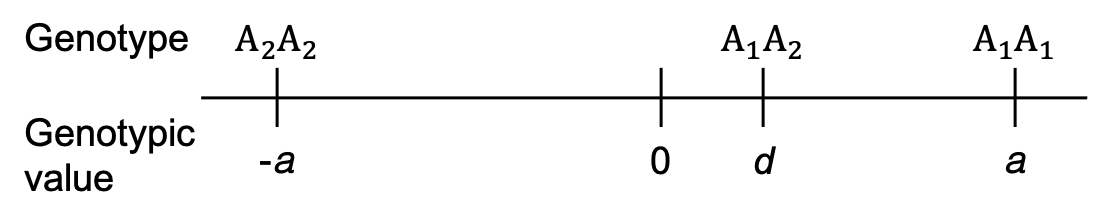
Arbitrarily assigned genotypic values (i.e., trait means per genotype class)

Arbitrarily assigned genotypic values (i.e., trait means per genotype class)
Variance components
Graphical representation of genotypic values (closed circles) at two biallelic loci A and B. The horizontal scale show the number of A1 alleles in the genotype. The different genotypes at locus B are depicted in different colors. For each of the locus B genotypes, the linear regression line between the number of A1 allele and the genotypic value is drawn.
Reproduction of the Figure 7.2 of Falconer and Mackay (1996). Graphical representation of genotypic (closed blue circles) and breeding (open blue circles) values,
of the genotypes for a locus with two alleles A1 and A2 at freqencies p and 1-p.
Horizontal scale: number of A1 allele in the genotype. Vertical scales of values are : on the left-arbitrarily assigned values (see figure at the top of the page);
on right-deviation from the population mean (black cross). Each point size is weighted by its genotype frequency. A linear regression line between the number of A1 allele and the genotypic values is fitted by weighted least square.
Additive (VA), additive-by-additive (VAA) and proportion of genotypic variance explained by additive variance (VA/VG) as a function of the allele frequencies p and q. Current setting of p and q is depicted with a white cross.
Distributions of additive (VA), dominance (VD) and total (VG) genetic variance on the left, and proportion of genotypic variance explained by additive variance (VA/VG) as a function of the allele frequency p on the right. The current user input allele frequency p is depicted by a vertical red solid line in the left panel and by a red cross in the right panel.